Grove Lab News
Viro3D Joins the AlphaFold Database
Feb 2026: Our Viro3D viral protein structure predictions have been incorporated into the EMBL-EBI AlphaFold Database as a community dataset. The AlphaFold Database now hosts our collection of 85,000+ viral protein structure predictions spanning 4,400 human and animal viruses, making them accessible through the world’s leading protein structure resource. Read the EMBL-EBI announcement.
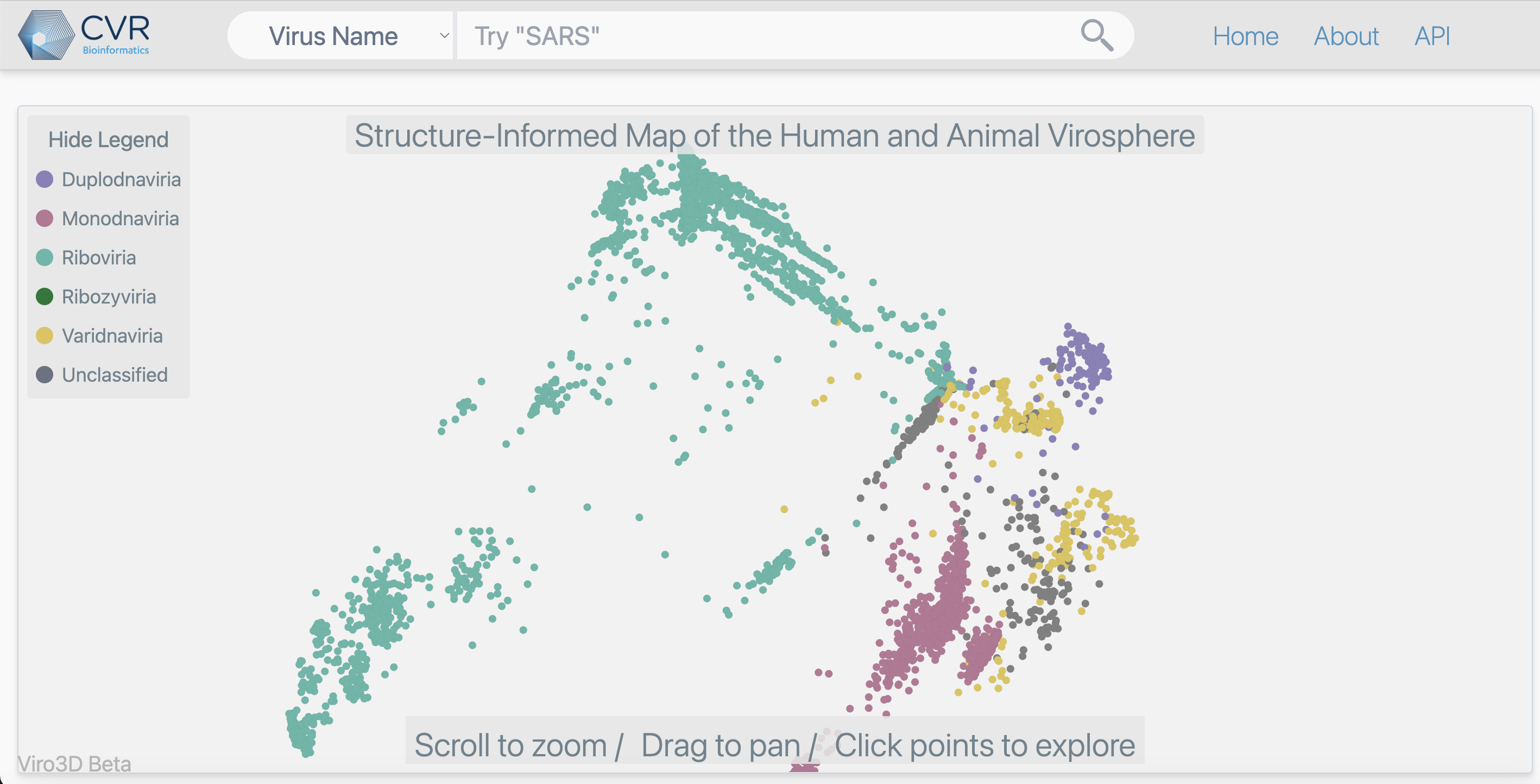
Launch of Viro3D Database
Oct 2025: We’re excited to announce the publication of "Viro3D: a comprehensive database of virus protein structure predictions". Using cutting-edge AI tools (AlphaFold2-ColabFold and ESMFold), we’ve predicted structures for 85,000 proteins from 4,400 human and animal viruses, expanding the structural coverage of viral proteins by 30 times compared to experimental structures. This was a team-CVR collaboration with our colleagues Ulad Litvin, Spyros Lytras, Alexander Jack, David Robertson, and Joseph Hughes. We’ve made all these structures freely available through our new interactive resource: https://viro3d.cvr.gla.ac.uk/

Featured on the BBC
Sept 2025: Our work was featured in the BBC documentary Disease X: Hunting the Next Pandemic, alongside presenter Dr. Chris van Tulleken. The Horizon documentary explores the science of pandemic preparedness and the hunt for the pathogen that could trigger the next global outbreak.

Emerging Viruses Symposium, Oxford
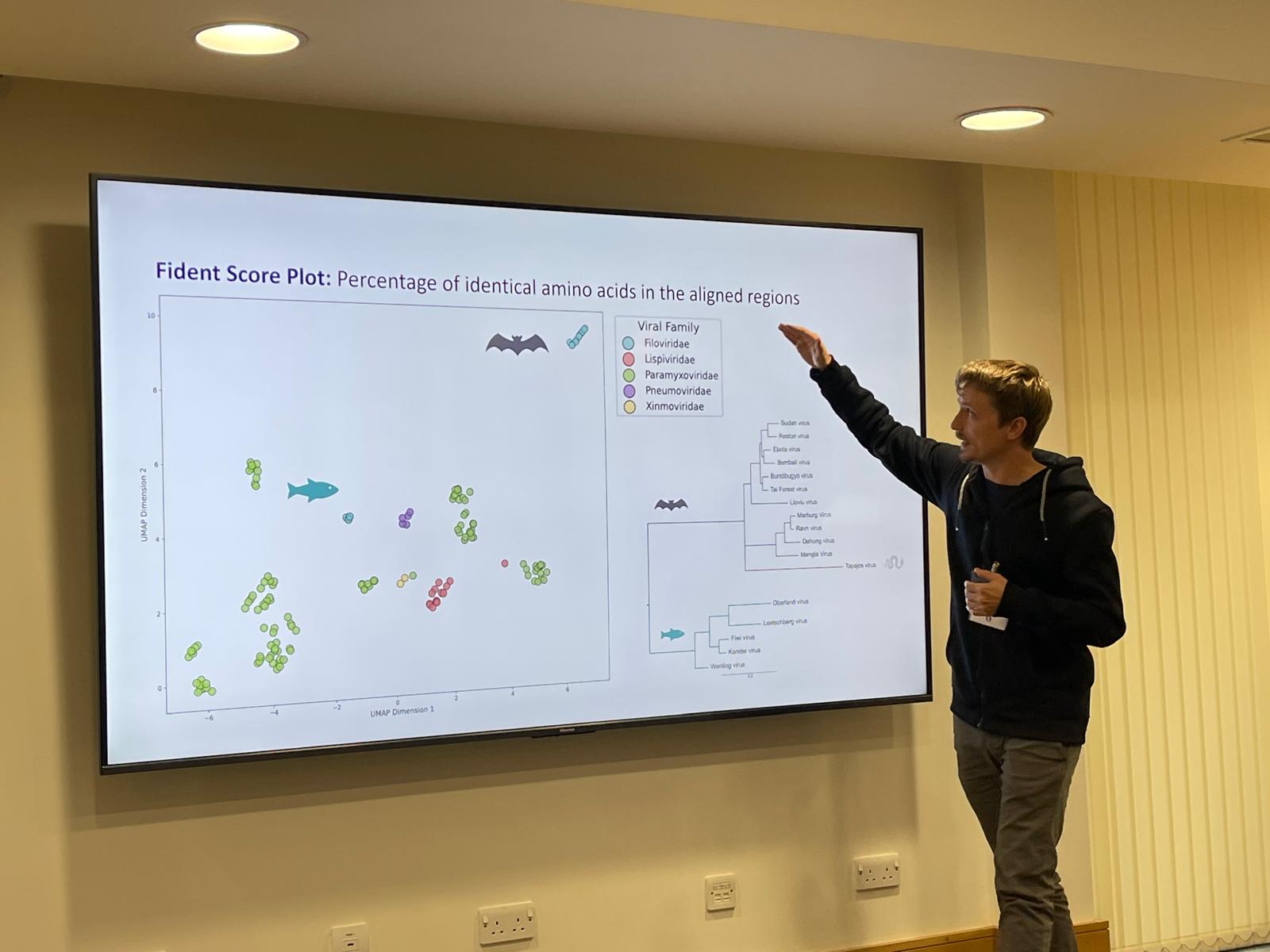
Sept 2025: Diego Cantoni presented our talk "Emerging AIs vs Emerging Viruses" at the Emerging Viruses Symposium 2025, University of Oxford. The talk showcased how state-of-the-art AI tools can be applied to newly emerging viruses, highlighting their potential to streamline research and accelerate pandemic preparedness.

AI & Pandemic Preparedness Roundtable
Aug 2025: Joe co-chaired a roundtable with UKHSA, bringing together experts from across government, academia, and public health to explore how biological AI can strengthen pandemic preparedness. Discussions centred on two themes: using machine learning for genomic surveillance and zoonotic risk assessment, and harnessing protein structure prediction tools to build libraries of therapeutic targets and vaccine antigens ahead of any future outbreak — a strategy of pre-emptive, knowledge stockpiling.

On the Road
May 2025: We recently had the opportunity to share our research at the Australian Virology Society, German Society for Virology, and European Congress of Virology meetings. It was a real privilege to connect with these vibrant and impressive scientific communities.





Cakes!
March 2025: We’d like to take a moment to celebrate the cakes that Sarah has made for various lab events. Not only are they delicious but also expertly decorated with lab memes.
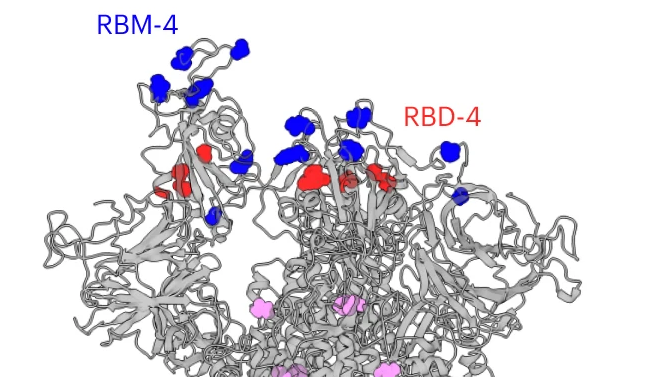
New Publication on SARS-CoV-2 Evolution
Jan 2025: Diego Cantoni from our team contributed to an exciting study, in collaboration with colleagues at the CVR, tracking how SARS-CoV-2 spike proteins evolved during the pandemic. The work, led by Wil Furnon, Vanessa Cowton, and Giuditta De Lorenzo, used viruses with different spike variants representing SARS-CoV-2 evolution from 2020-2024.

RSE Small Grant Award
Sept 2024: Diego Cantoni secured a Royal Society of Edinburgh Small Grant Award to characterise the elusive secreted glycoproteins of filoviruses. Congratulations Diego!
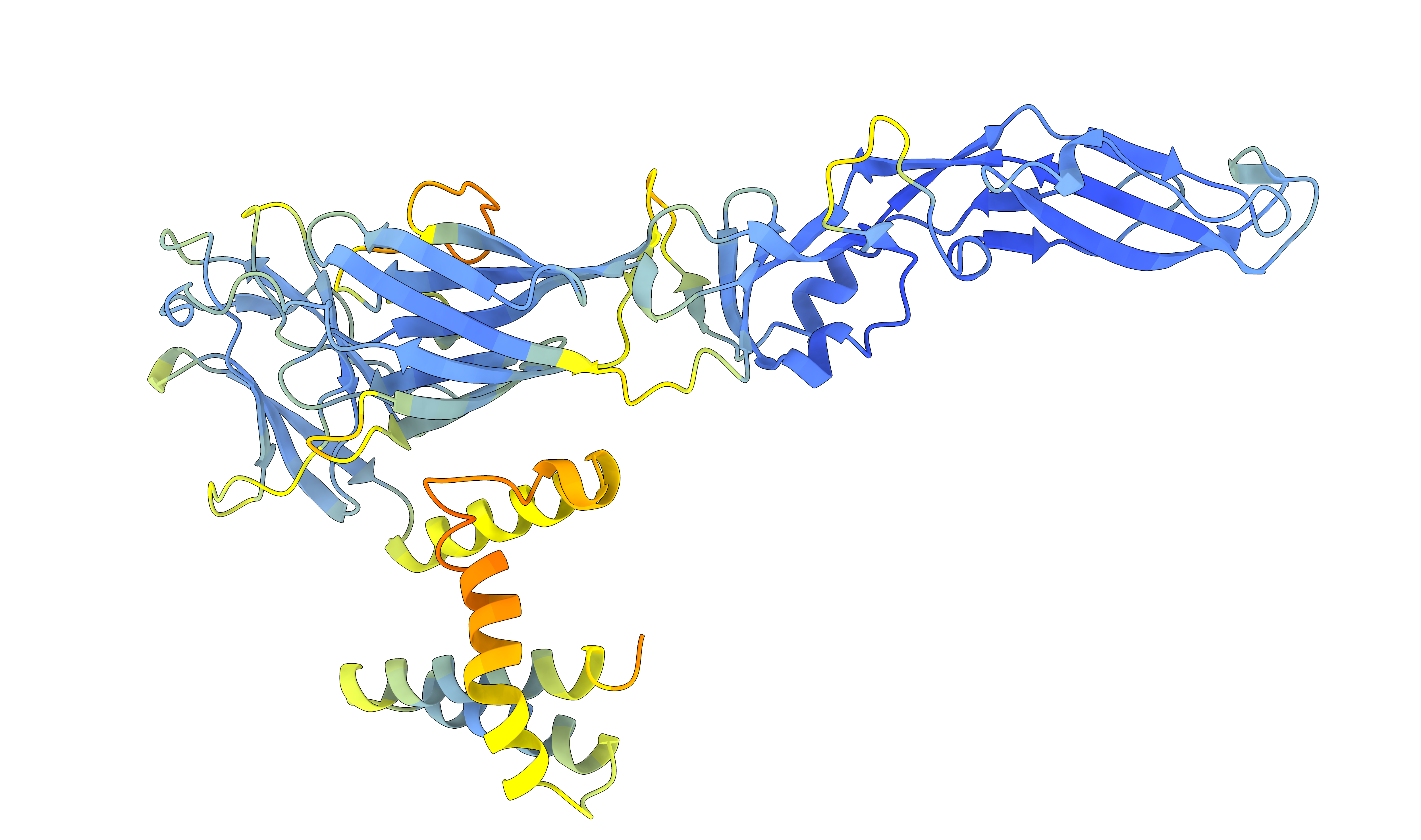
New Publication on Flaviviridae Glycoproteins
Sept 2024: We’re pleased to share our new study published in Nature which maps glycoprotein structures across the entire Flaviviridae family. By combining phylogenetic analyses with protein structure prediction, we’ve surveyed glycoproteins in this diverse virus family that includes important pathogens such as hepatitis C, dengue and Zika viruses. Our findings reveal class II fusion systems in most species, while highlighting structurally distinct E1E2 glycoproteins in hepaciviruses, pegiviruses and pestiviruses. This work, in collaboration with colleagues at the University of Sydney, provides insights into viral fusion mechanisms and evolutionary history that have shaped the virology and ecology of these significant pathogens.